Part 2: Antigen Selection Pipeline
2.3. Pipeline Recap
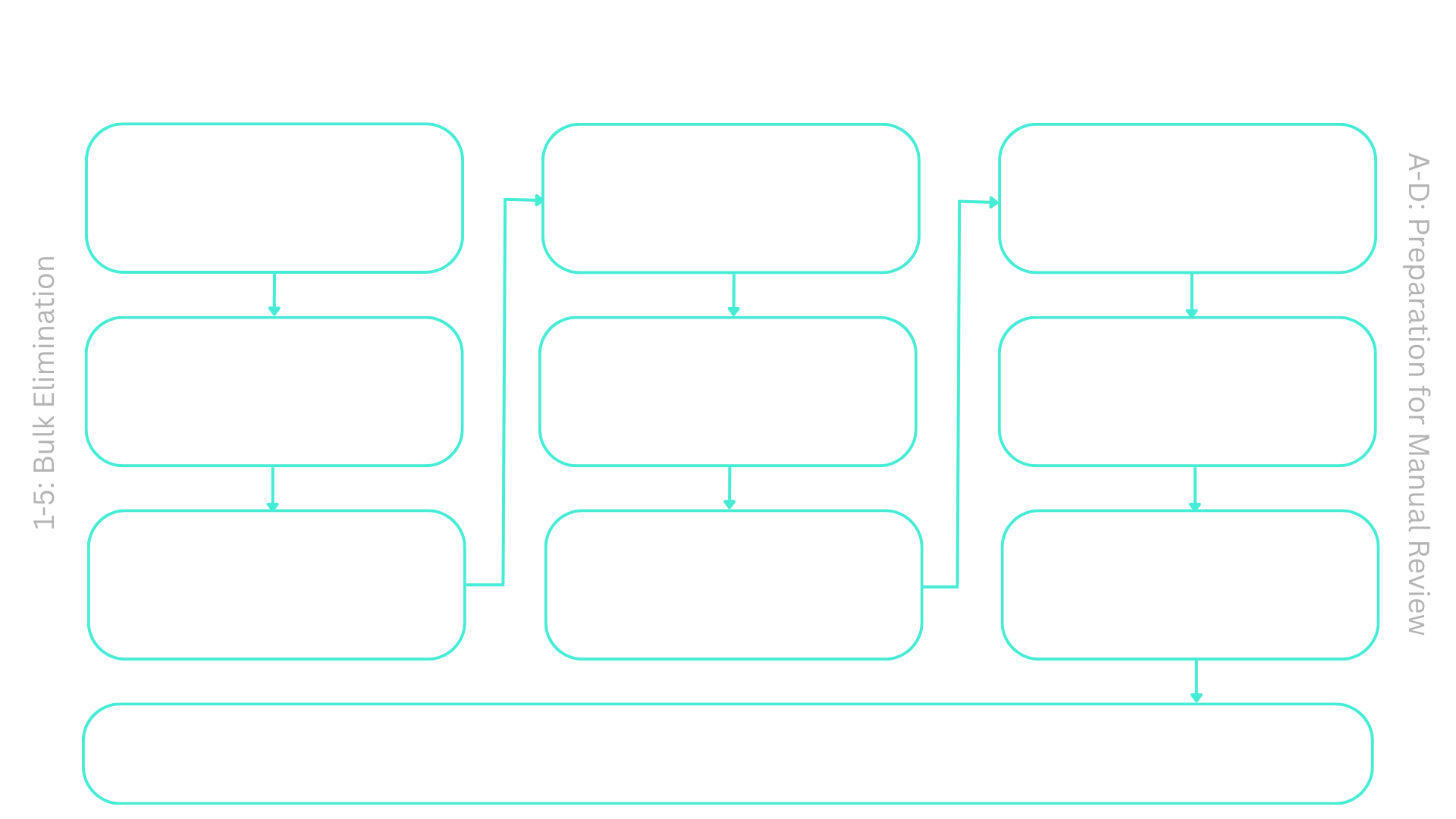
This marks the end of the bulk elimination phase of the pipeline, which screens and suggests a few promising candidates. As we’ve just seen, the pipeline gathers a lot of initial data (step 1), then removes proteins with a low likelihood of being good antigens using several filters. The filters can be built by first thinking about the attributes of a good antigen. In step 2, all proteins not shared across most strains were removed, as a good antigen would be vital and conserved against A. baumannii subspecies. In step 3, we removed proteins that were too similar to human or gut microbiome proteins to minimize the risk that the vaccine could cause major harm to human subjects. In step 4, we used the fact that an antigen is readily accessible and filtered out proteins found only inside bacterial cells. Lastly, in step 5, we ensure that suggested candidates are sufficiently similar to virulence proteins that are also essential for bacterial function, thereby reducing the risk of developing resistance to antibodies targeting the antigens. Overall, filters are somewhat lax in accounting for the potential errors a bioinformatics tool may produce - still, by getting proteins through multiple filters, a manageable list of suggestions can be created. With some manual curation, some of the suggested candidates can then be tested in a lab.